Selected Publications
Nature Machine Intelligence, 2025
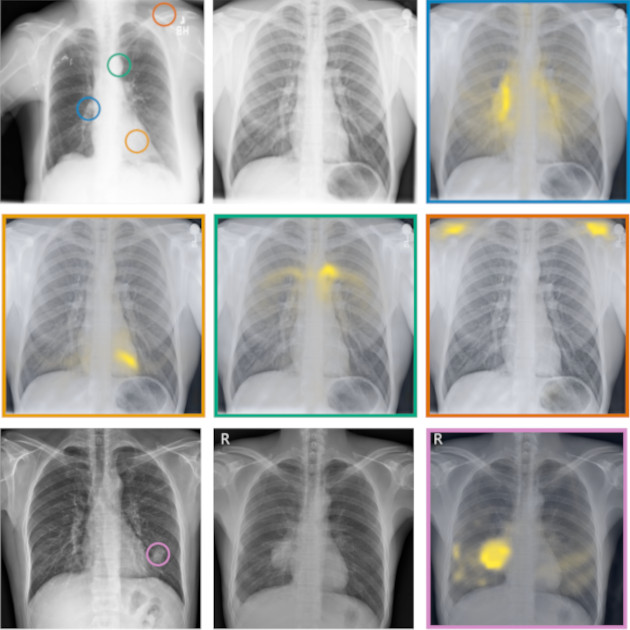
Language-supervised pretraining has proven to be a valuable method for extracting semantically meaningful features from images, serving as a foundational element in multimodal systems within the computer vision and medical imaging domains. However, the computed features are limited by the information contained in the text, which is particularly problematic in medical imaging, in which the findings described by radiologists focus on specific observations. This challenge is compounded by the scarcity of paired imaging–text data due to concerns over the leakage of personal health information. In this work, we fundamentally challenge the prevailing reliance on language supervision for learning general-purpose biomedical imaging encoders. We introduce RAD-DINO, a biomedical image encoder pretrained solely on unimodal biomedical imaging data that obtains similar or greater performance than state-of-the …
arXiv preprint arXiv:2406.04449, 2024
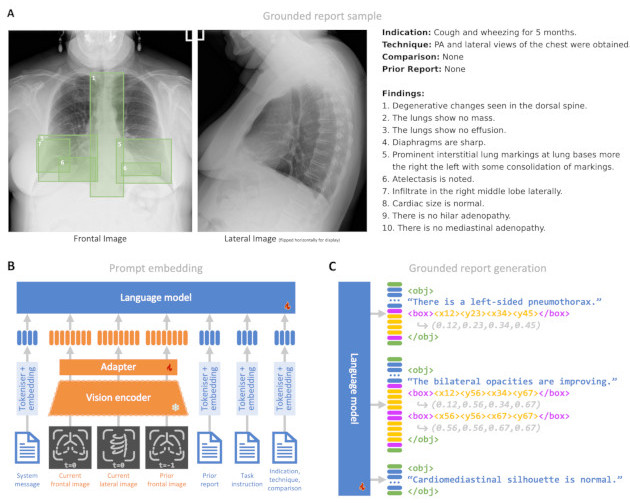
Radiology reporting is a complex task requiring detailed medical image understanding and precise language generation, for which generative multimodal models offer a promising solution. However, to impact clinical practice, models must achieve a high level of both verifiable performance and utility. We augment the utility of automated report generation by incorporating localisation of individual findings on the image - a task we call grounded report generation - and enhance performance by incorporating realistic reporting context as inputs. We design a novel evaluation framework (RadFact) leveraging the logical inference capabilities of large language models (LLMs) to quantify report correctness and completeness at the level of individual sentences, while supporting the new task of grounded reporting. We develop MAIRA-2, a large radiology-specific multimodal model designed to generate chest X-ray reports with and without grounding. MAIRA-2 achieves state of the art on existing report generation benchmarks and establishes the novel task of grounded report generation.
European Conference on Computer Vision (ECCV), 2024
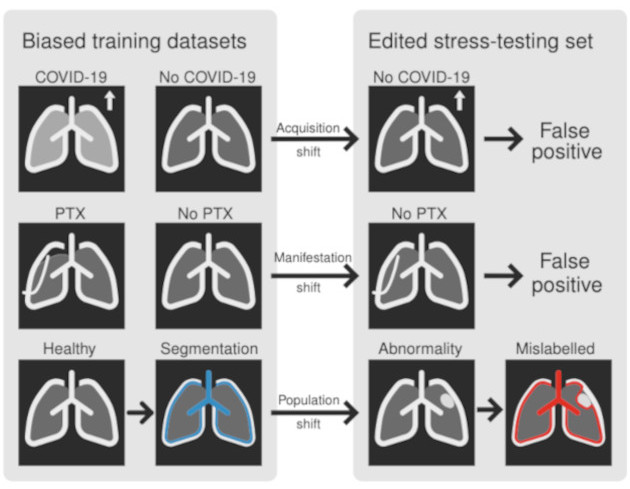
Biomedical imaging datasets are often small and biased, meaning that real-world performance of predictive models can be substantially lower than expected from internal testing. This work proposes using generative image editing to simulate dataset shifts and diagnose failure modes of biomedical vision models; this can be used in advance of deployment to assess readiness, potentially reducing cost and patient harm. Existing editing methods can produce undesirable changes, with spurious correlations learned due to the co-occurrence of disease and treatment interventions, limiting practical applicability. To address this, we train a text-to-image diffusion model on multiple chest X-ray datasets and introduce a new editing method, RadEdit, that uses multiple image masks, if present, to constrain changes and ensure consistency in the edited images, minimising bias. We consider three types of dataset shifts …
Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2023
Self-supervised learning in vision--language processing (VLP) exploits semantic alignment between imaging and text modalities. Prior work in biomedical VLP has mostly relied on the alignment of single image and report pairs even though clinical notes commonly refer to prior images. This does not only introduce poor alignment between the modalities but also a missed opportunity to exploit rich self-supervision through existing temporal content in the data. In this work, we explicitly account for prior images and reports when available during both training and fine-tuning. Our approach, named BioViL-T, uses a CNN--Transformer hybrid multi-image encoder trained jointly with a text model. It is designed to be versatile to arising challenges such as pose variations and missing input images across time. The resulting model excels on downstream tasks both in single-and multi-image setups, achieving state-of-the-art (SOTA) performance on (I) progression classification,(II) phrase grounding, and (III) report generation, whilst offering consistent improvements on disease classification and sentence-similarity tasks. We release a novel multi-modal temporal benchmark dataset, CXR-T, to quantify the quality of vision--language representations in terms of temporal semantics. Our experimental results show the significant advantages of incorporating prior images and reports to make most use of the data.
Computer Methods and Programs in Biomedicine, 2021
Background and Objective. Processing of medical images such as MRI or CT presents different challenges compared to RGB images typically used in computer vision. These include a lack of labels for large datasets, high computational costs, and the need of metadata to describe the physical properties of voxels. Data augmentation is used to artificially increase the size of the training datasets. Training with image subvolumes or patches decreases the need for computational power. Spatial metadata needs to be carefully taken into account in order to ensure a correct alignment and orientation of volumes. We present TorchIO, an open-source Python library to enable efficient loading, preprocessing, augmentation and patch-based sampling of medical images for deep learning. TorchIO follows the style of PyTorch and integrates standard medical image processing libraries to efficiently process images during …
Medical Image Analysis, 2024
The lack of annotated datasets is a major bottleneck for training new task-specific supervised machine learning models, considering that manual annotation is extremely expensive and time-consuming. To address this problem, we present MONAI Label, a free and open-source framework that facilitates the development of applications based on artificial intelligence (AI) models that aim at reducing the time required to annotate radiology datasets. Through MONAI Label, researchers can develop AI annotation applications focusing on their domain of expertise. It allows researchers to readily deploy their apps as services, which can be made available to clinicians via their preferred user interface. Currently, MONAI Label readily supports locally installed (3D Slicer) and web-based (OHIF) frontends and offers two active learning strategies to facilitate and speed up the training of segmentation algorithms. MONAI Label …